| |
News
Our friends (who are always very supportive of the AFND website, Jill A. Hollenbach @UCSF, Steven J. Mack @UCSF and Martin Maiers @NMDP) have formed
the COVID-19 HLA & Immunogenetics Consortium to unite the global community of HLA and immunogenetics experts and leaders in support of these efforts.
They invite you to share your expertise as they launch this endeavour by joining the consortium. A project for the next international Histocompatibility
Workshop is also planned.
Introduction
The Allele Frequency Net Database (AFND) provides the scientific community with a freely available repository for the storage
of immune gene frequencies in different worldwide populations. Users can contribute the results of their work into one common database
and can perform database searches on information already available. We have currently collected data in allele,
haplotype and genotype format. However, the success of this website will depend on you to contribute your data.
Does any of this interest you?
How good are the data sets in AFND?
We have classified HLA data sets into "Gold-Silver-Bronce" standards. If you wish to see how many populations are classified as Gold,
click here. 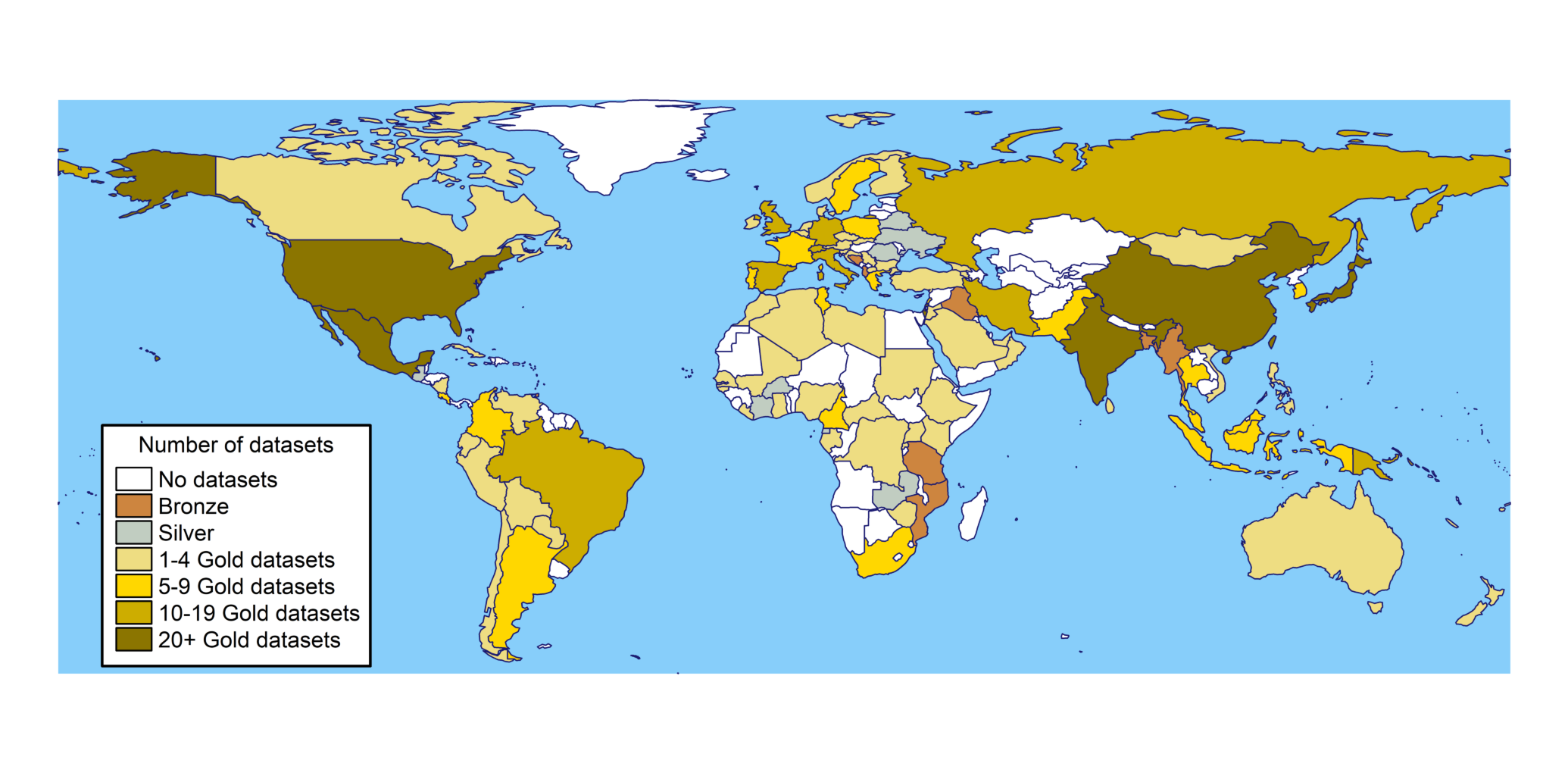
Please cite this website using our last publication:
Allele frequency net database (AFND) 2020 update: gold-standard data classification, open access genotype data and new query tools.
Gonzalez-Galarza FF, McCabe A, Santos EJ, Jones J, Takeshita LY, Ortega-Rivera ND, Del Cid-Pavon GM, Ramsbottom K, Ghattaoraya GS, Alfirevic A, Middleton D and Jones AR Nucleic Acid Research 2020, 48, D783-8.
[Full Text]
We recommend you to also cite the original publication report of the data in your references.
Database information
| Polymorphic Region |
Population Studies |
Gene/Allele Data |
Haplotype Data |
Genotype Data |
| HLA | 1324 | 1306 | 662 | - |
| KIR | 293 | 292 | - | 192 |
| Cytokine | 125 | 125 | - | - |
| MIC | 64 | 64 | 23 | - |
| Totals |
1,806 |
1,787 |
685 |
192 |
The current number of frequencies stored in our database is:
156,049 (HLA),
6,731 (KIR),
4,376 (Cytokine) and
877 (MIC) from
14,264,556 individuals.
We have updated the website with the new IMGT/HLA nomenclature
guidelines.
IMGT/HLA
last Update: 3.37.0, 10 July, 2019. |
IPD-KIR
last Update: 2.6.0, 17 February, 2015.
|
|
|
Login
You are logged as: guest
Click here to use your account
SUBMIT YOUR DATA
Sponsors
Most popular searches
Latest developments
|